RNA polyadenylation and its consequences in prokaryotes | Philosophical Transactions of the Royal Society B: Biological Sciences
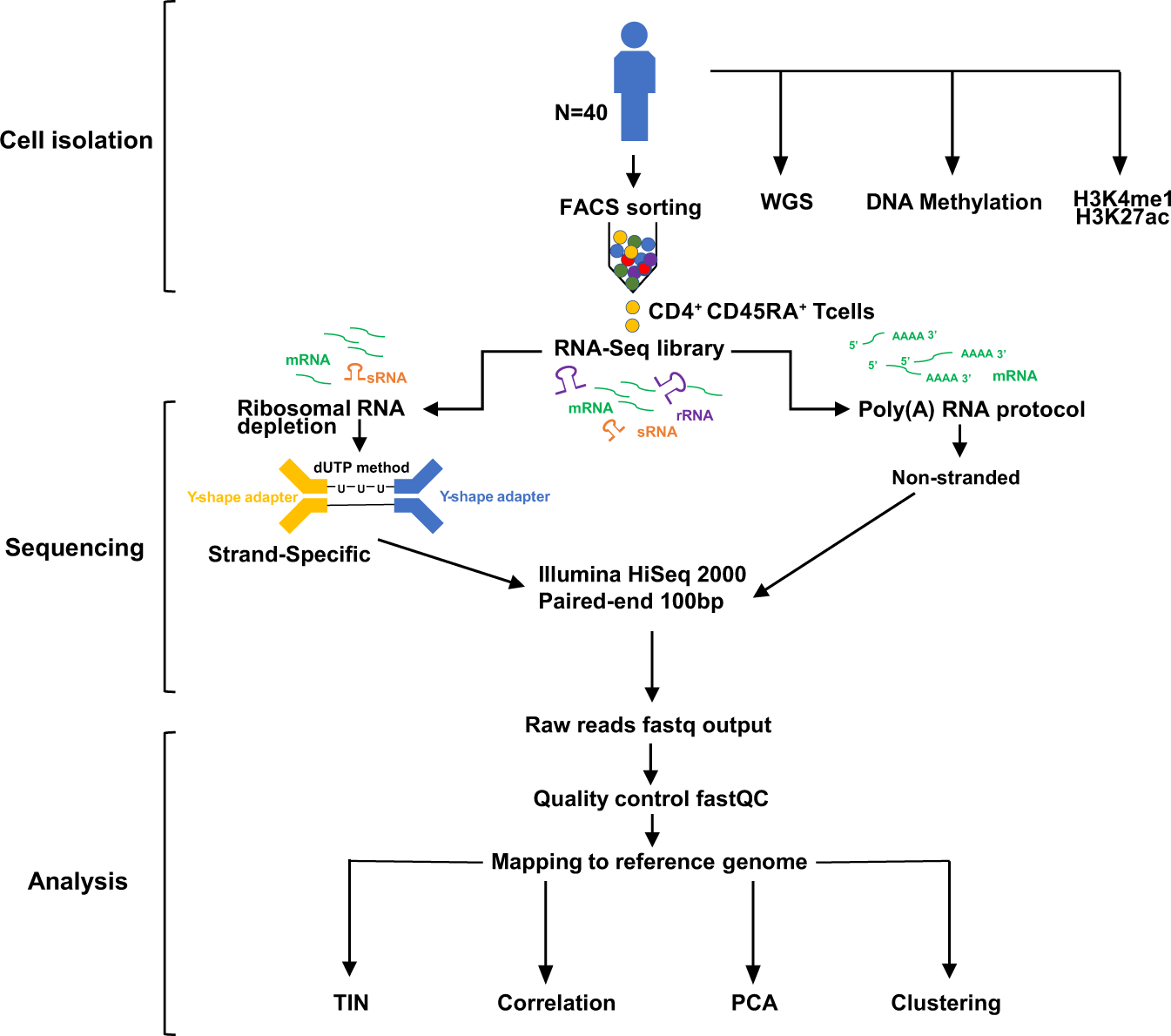
Paired rRNA-depleted and polyA-selected RNA sequencing data and supporting multi-omics data from human T cells | Scientific Data
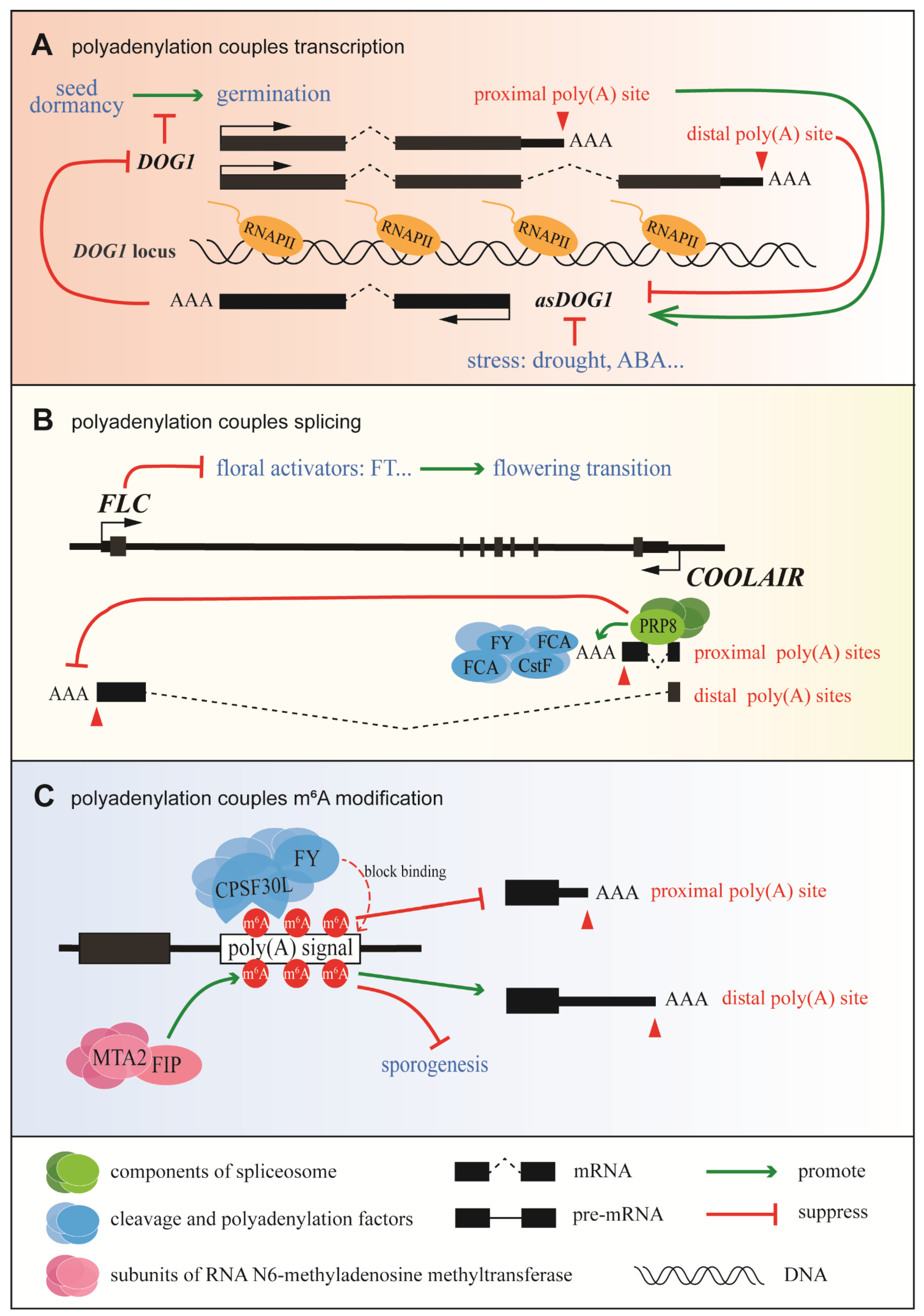
IJMS | Free Full-Text | Co-Transcriptional RNA Processing in Plants: Exploring from the Perspective of Polyadenylation | HTML
Characterization of the Role of Hexamer AGUAAA and Poly(A) Tail in Coronavirus Polyadenylation | PLOS ONE
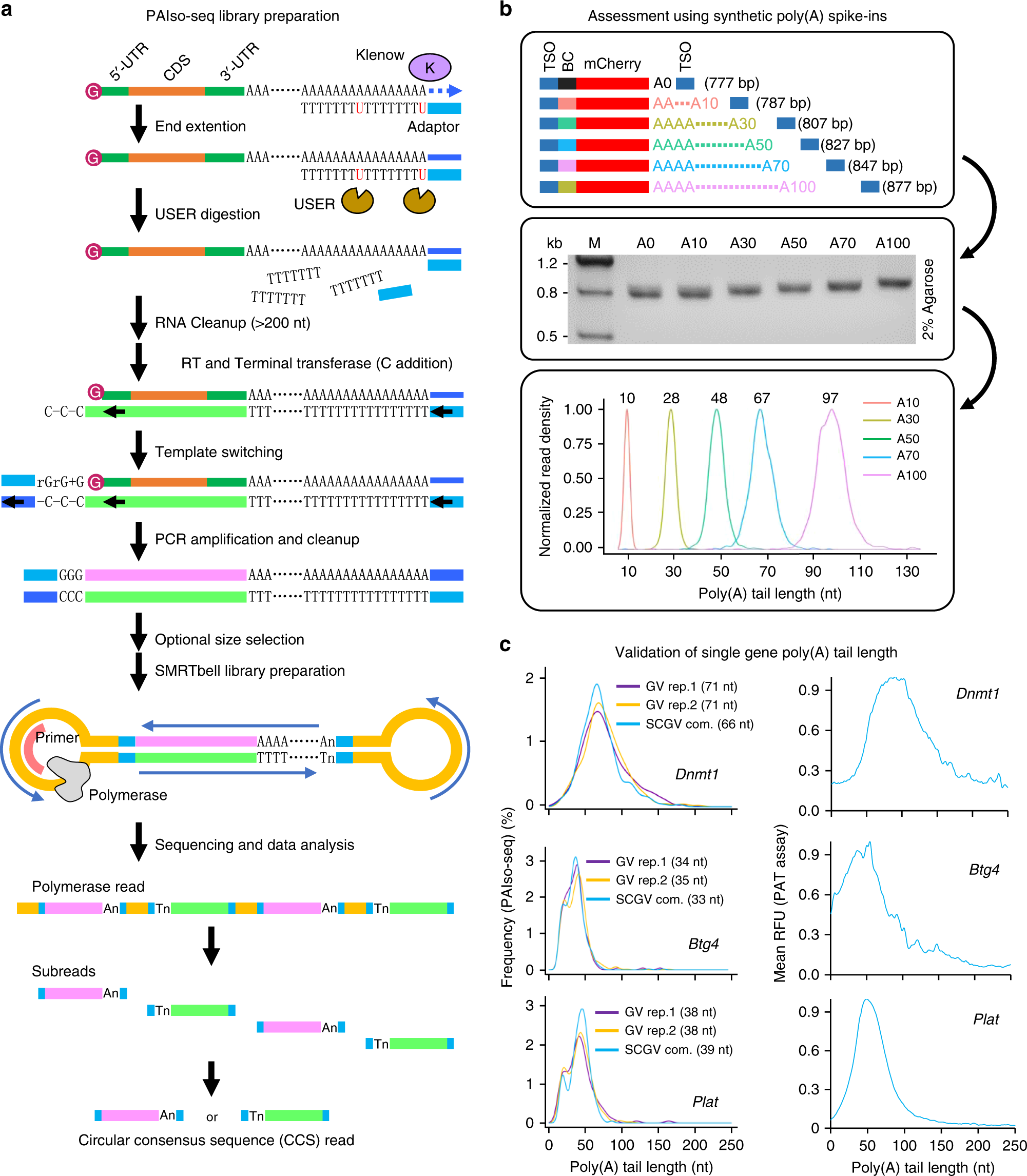
Poly(A) inclusive RNA isoform sequencing (PAIso−seq) reveals wide-spread non-adenosine residues within RNA poly(A) tails | Nature Communications
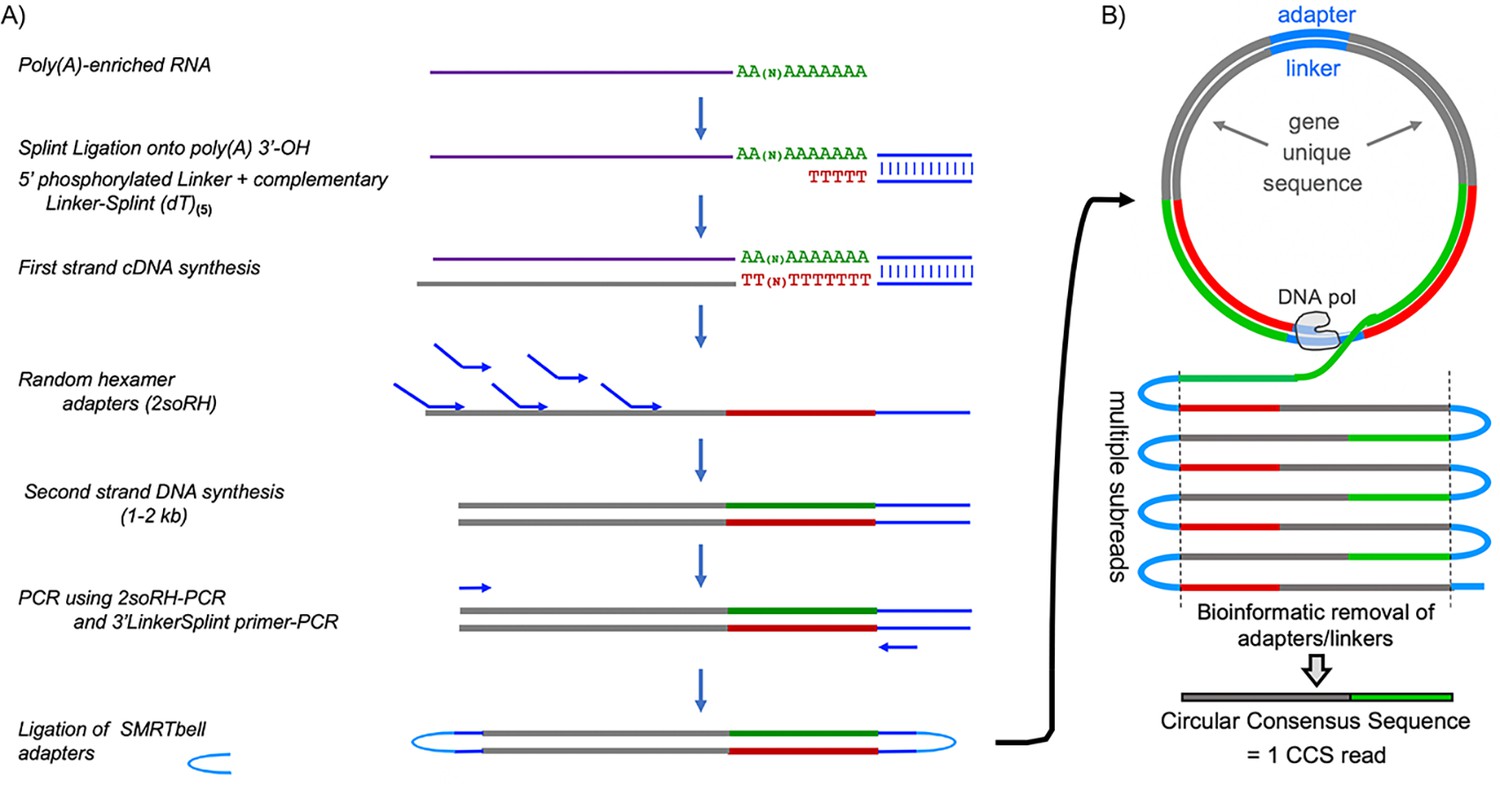
Single molecule poly(A) tail-seq shows LARP4 opposes deadenylation throughout mRNA lifespan with most impact on short tails | eLife
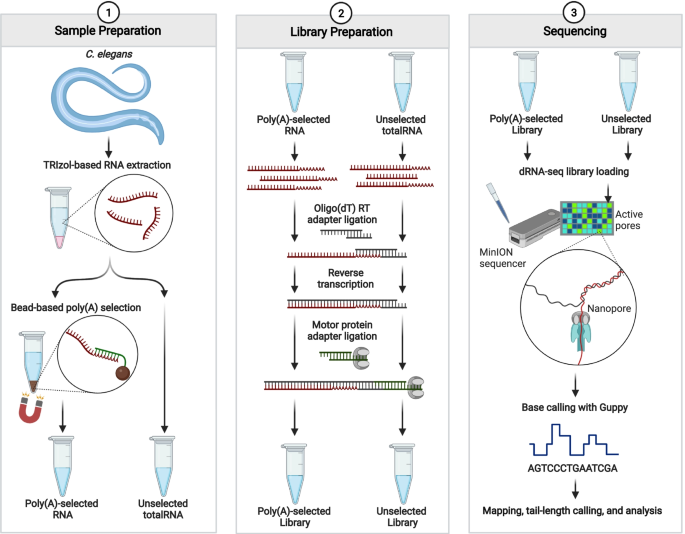
Poly(a) selection introduces bias and undue noise in direct RNA-sequencing | BMC Genomics | Full Text

The intrinsic structure of poly(A) RNA determines the specificity of Pan2 and Caf1 deadenylases | Nature Structural & Molecular Biology

A multiplex RNA-seq strategy to profile poly(A+) RNA: Application to analysis of transcription response and 3′ end formation - ScienceDirect

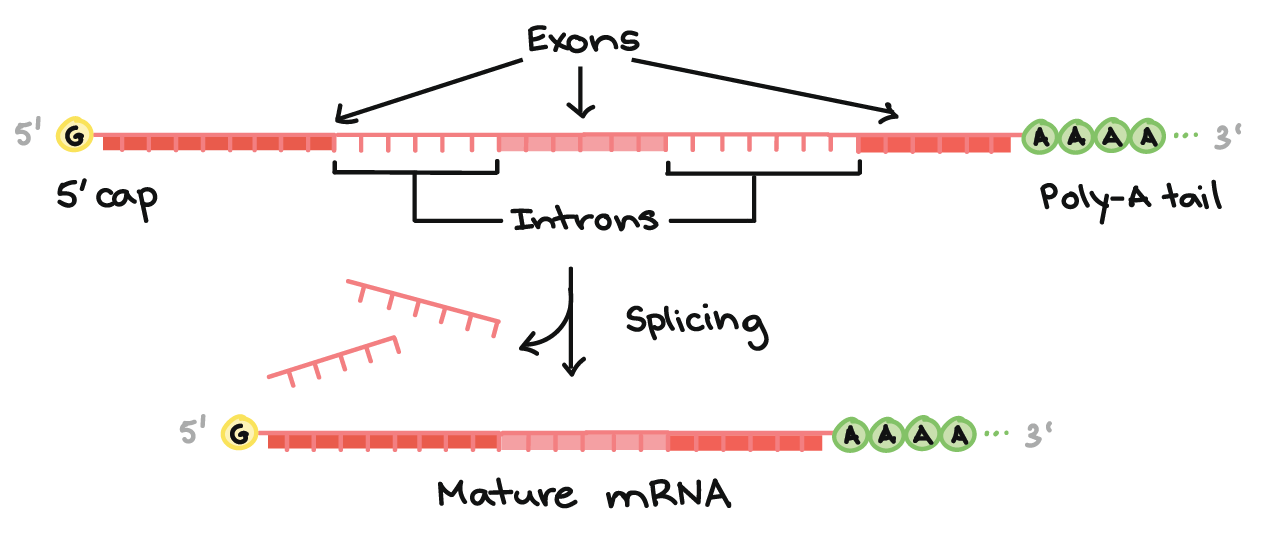
RNA processing begins with the addition of a 5'-cap and a 3'-poly(rA)... | Download Scientific Diagram
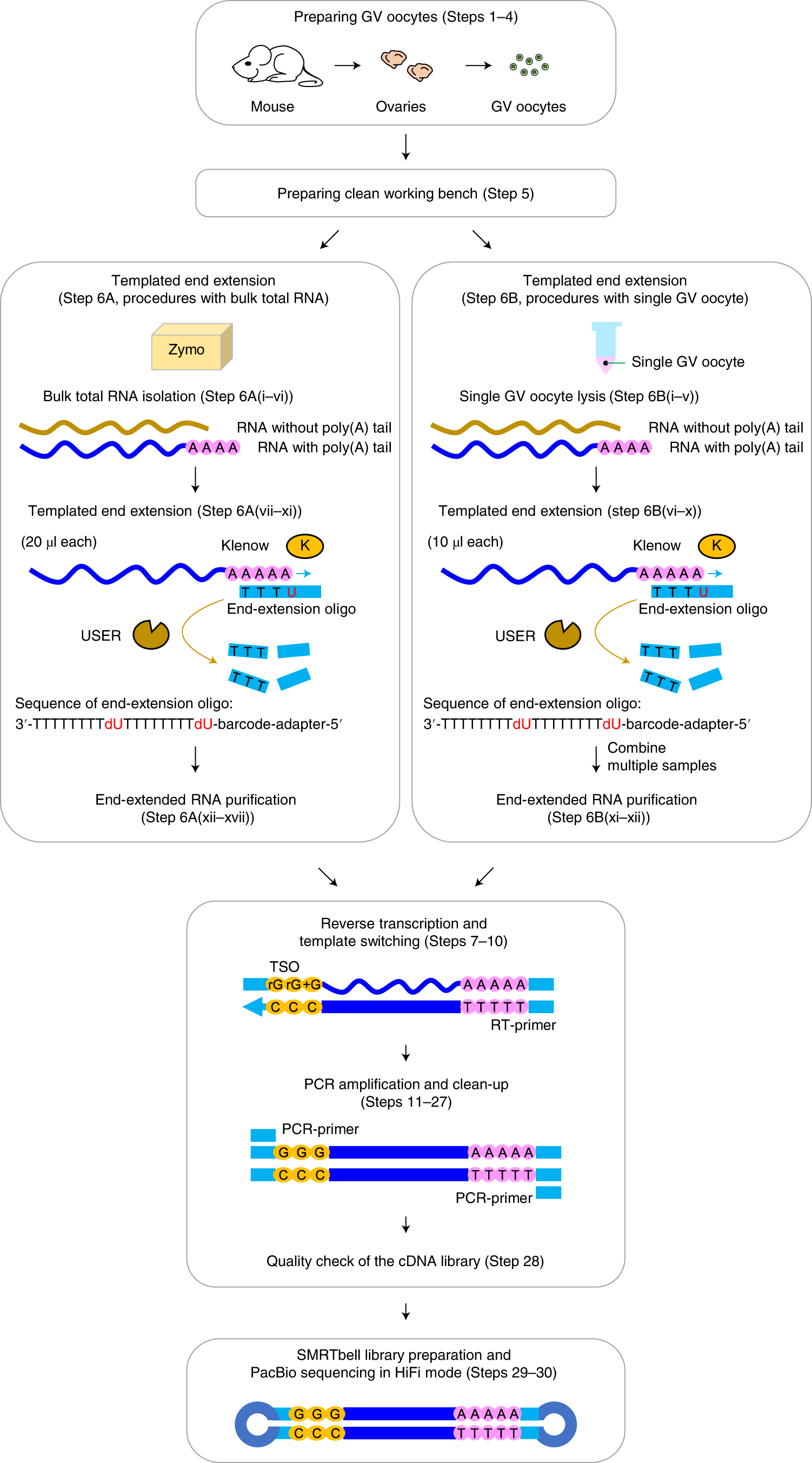
Transcriptome-wide measurement of poly(A) tail length and composition at subnanogram total RNA sensitivity by PAIso-seq | Nature Protocols

Influenza A Virus NS1 Protein Binds as a Dimer to RNA-Free PABP1 but Not to the PABP1·Poly(A) RNA Complex | Biochemistry